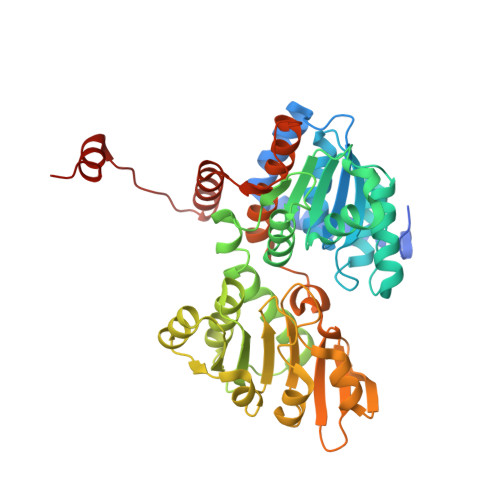
Structure, function and substrate preferences of archaeal S-adenosyl-L-homocysteine hydrolases.
Koeppl, L.H., Popadic, D., Saleem-Batcha, R., Germer, P., Andexer, J.N.(2024) Commun Biol 7: 380-380
- PubMed: 38548921
- DOI: https://doi.org/10.1038/s42003-024-06078-9
- Primary Citation of Related Structures:
7R37, 7R38, 7R39, 7R3A, 8COD, 8QNO - PubMed Abstract:
S-Adenosyl-L-homocysteine hydrolase (SAHH) reversibly cleaves S-adenosyl-L-homocysteine, the product of S-adenosyl-L-methionine-dependent methylation reactions. The conversion of S-adenosyl-L-homocysteine into adenosine and L-homocysteine plays an important role in the regulation of the methyl cycle. An alternative metabolic route for S-adenosyl-L-methionine regeneration in the extremophiles Methanocaldococcus jannaschii and Thermotoga maritima has been identified, featuring the deamination of S-adenosyl-L-homocysteine to S-inosyl-L-homocysteine. Herein, we report the structural characterisation of different archaeal SAHHs together with a biochemical analysis of various SAHHs from all three domains of life. Homologues deriving from the Euryarchaeota phylum show a higher conversion rate with S-inosyl-L-homocysteine compared to S-adenosyl-L-homocysteine. Crystal structures of SAHH originating from Pyrococcus furiosus in complex with SLH and inosine as ligands, show architectural flexibility in the active site and offer deeper insights into the binding mode of hypoxanthine-containing substrates. Altogether, the findings of our study support the understanding of an alternative metabolic route for S-adenosyl-L-methionine and offer insights into the evolutionary progression and diversification of SAHHs involved in methyl and purine salvage pathways.
- Institute of Pharmaceutical Sciences, University of Freiburg, Albertstr. 25, 79104, Freiburg, Germany.
Organizational Affiliation: