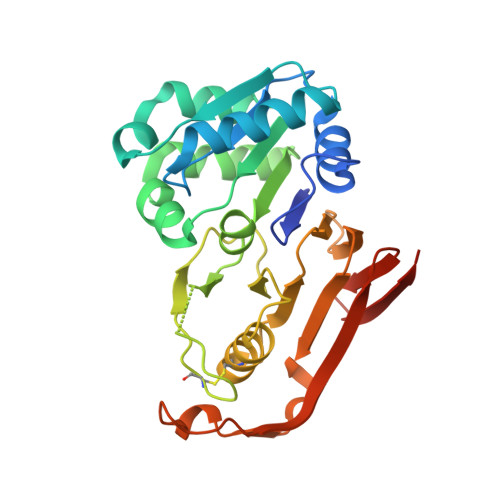
The structure of cyanophycinase in complex with a cyanophycin degradation intermediate.
Sharon, I., Grogg, M., Hilvert, D., Schmeing, T.M.(2022) Biochim Biophys Acta Gen Subj 1866: 130217-130217
- PubMed: 35905922
- DOI: https://doi.org/10.1016/j.bbagen.2022.130217
- Primary Citation of Related Structures:
7UQV, 7UQW - PubMed Abstract:
Cyanophycinases are serine protease family enzymes which are required for the metabolism of cyanophycin, the natural polymer multi-L-arginyl-poly(L-aspartic acid). Cyanophycinases degrade cyanophycin to β-Asp-Arg dipeptides, which enables use of this important store of fixed nitrogen. We used genetic code expansion to incorporate diaminopropionic acid into cyanophycinase in place of the active site serine, and determined a high-resolution structure of the covalent acyl-enzyme intermediate resulting from attack of cyanophycinase on a short cyanophycin segment. The structure indicates that cyanophycin dipeptide residues P1 and P1' bind shallow pockets adjacent to the catalytic residues. We observe many cyanophycinase - P1 dipeptide interactions in the co-complex structure. Calorimetry measurements show that at least two cyanophycin dipeptides are needed for high affinity binding to cyanophycinase. We also characterized a putative cyanophycinase which we found to be structurally very similar but that shows no activity and could not be activated by mutation of its active site. Despite its peptidic structure, cyanophycin is resistant to degradation by peptidases and other proteases. Our results help show how cyanophycinase can specifically bind and degrade this important polymer.
- Department of Biochemistry and Centre de recherche en biologie structurale, McGill University, Montréal, QC H3G 0B1, Canada.
Organizational Affiliation: