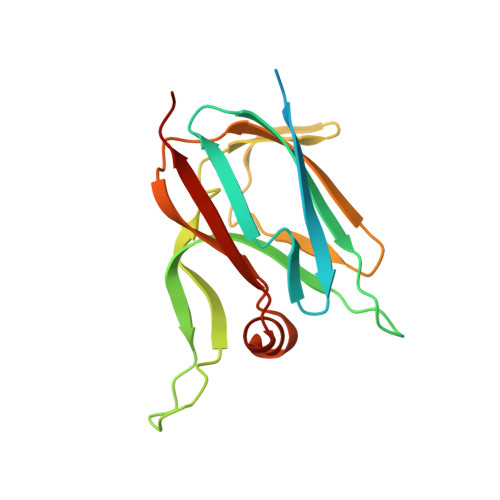
Beyond the coupled distortion model: structural analysis of the single domain cupredoxin AcoP, a green mononuclear copper centre with original features.
Roger, M., Leone, P., Blackburn, N.J., Horrell, S., Chicano, T.M., Biaso, F., Giudici-Orticoni, M.T., Abriata, L.A., Hura, G.L., Hough, M.A., Sciara, G., Ilbert, M.(2024) Dalton Trans 53: 1794-1808
- PubMed: 38170898
- DOI: https://doi.org/10.1039/d3dt03372d
- Primary Citation of Related Structures:
7Z3B, 7Z3F, 7Z3G, 7Z3I - PubMed Abstract:
Cupredoxins are widely occurring copper-binding proteins with a typical Greek-key beta barrel fold. They are generally described as electron carriers that rely on a T1 copper centre coordinated by four ligands provided by the folded polypeptide. The discovery of novel cupredoxins demonstrates the high diversity of this family, with variations in terms of copper-binding ligands, copper centre geometry, redox potential, as well as biological function. AcoP is a periplasmic cupredoxin belonging to the iron respiratory chain of the acidophilic bacterium Acidithiobacillus ferrooxidans. AcoP presents original features, including high resistance to acidic pH and a constrained green-type copper centre of high redox potential. To understand the unique properties of AcoP, we undertook structural and biophysical characterization of wild-type AcoP and of two Cu-ligand mutants (H166A and M171A). The crystallographic structures, including native reduced AcoP at 1.65 Å resolution, unveil a typical cupredoxin fold. The presence of extended loops, never observed in previously characterized cupredoxins, might account for the interaction of AcoP with physiological partners. The Cu-ligand distances, determined by both X-ray diffraction and EXAFS, show that the AcoP metal centre seems to present both T1 and T1.5 features, in turn suggesting that AcoP might not fit well to the coupled distortion model. The crystal structures of two AcoP mutants confirm that the active centre of AcoP is highly constrained. Comparative analysis with other cupredoxins of known structures, suggests that in AcoP the second coordination sphere might be an important determinant of active centre rigidity due to the presence of an extensive hydrogen bond network. Finally, we show that other cupredoxins do not perfectly follow the coupled distortion model as well, raising the suspicion that further alternative models to describe copper centre geometries need to be developed, while the importance of rack-induced contributions should not be underestimated.
- CNRS, Aix-Marseille University, Bioenergetic and Protein Engineering Laboratory, BIP UMR 7281, Mediterranean Institute of Microbiology, 13009 Marseille, France. giuliano.sciara@inrae.fr.
Organizational Affiliation: