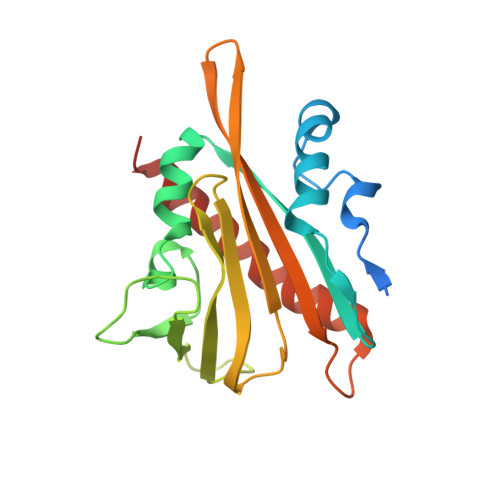
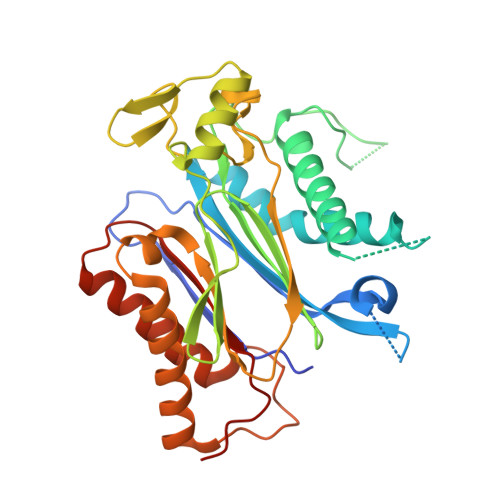
Structure-guided engineering of a receptor-agonist pair for inducible activation of the ABA adaptive response to drought.
Lozano-Juste, J., Infantes, L., Garcia-Maquilon, I., Ruiz-Partida, R., Merilo, E., Benavente, J.L., Velazquez-Campoy, A., Coego, A., Bono, M., Forment, J., Pampin, B., Destito, P., Monteiro, A., Rodriguez, R., Cruces, J., Rodriguez, P.L., Albert, A.(2023) Sci Adv 9: eade9948-eade9948
- PubMed: 36897942
- DOI: https://doi.org/10.1126/sciadv.ade9948
- Primary Citation of Related Structures:
6ZUC, 8AY3, 8AY6, 8AY7, 8AY8, 8AY9, 8AYA - PubMed Abstract:
Strategies to activate abscisic acid (ABA) receptors and boost ABA signaling by small molecules that act as ABA receptor agonists are promising biotechnological tools to enhance plant drought tolerance. Protein structures of crop ABA receptors might require modifications to improve recognition of chemical ligands, which in turn can be optimized by structural information. Through structure-based targeted design, we have combined chemical and genetic approaches to generate an ABA receptor agonist molecule (iSB09) and engineer a CsPYL1 ABA receptor, named CsPYL1 5m , which efficiently binds iSB09. This optimized receptor-agonist pair leads to activation of ABA signaling and marked drought tolerance. No constitutive activation of ABA signaling and hence growth penalty was observed in transformed Arabidopsis thaliana plants. Therefore, conditional and efficient activation of ABA signaling was achieved through a chemical-genetic orthogonal approach based on iterative cycles of ligand and receptor optimization driven by the structure of ternary receptor-ligand-phosphatase complexes.
Organizational Affiliation:
Instituto de Biología Molecular y Celular de Plantas, Consejo Superior de Investigaciones Científicas, Universidad Politécnica de Valencia, 46022 Valencia, Spain.