Venom protection by antibody from a snakebite hyperimmune subject
Glanville, J., Andrade, J., Bellin, M., Kim, S., Pletnev, S., Tsao, D., Verardi, R., Bedi, R., Friede, T., Tully, E., Zhang, B., Bylund, T., Liu, T., Kwong, P.D.To be published.
Experimental Data Snapshot
Starting Model: in silico
View more details
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Centi-LNX-D09 Fab light chain | 214 | Homo sapiens | Mutation(s): 0 | 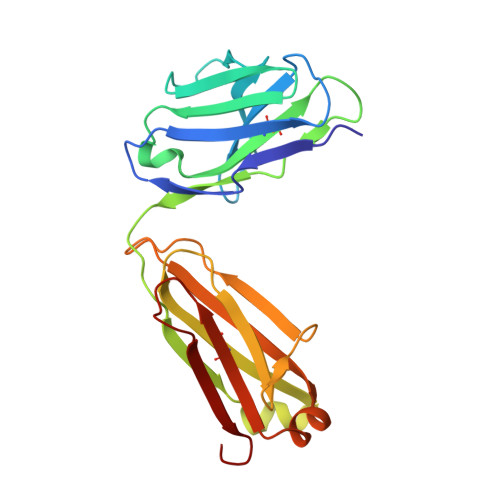 | |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Centi-LNX-D09 Fab heavy chain | 237 | Homo sapiens | Mutation(s): 0 | 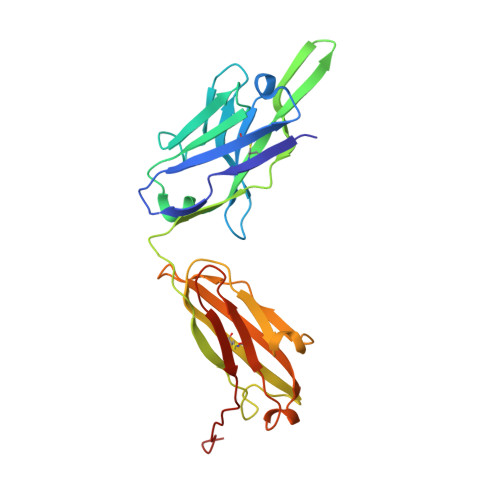 | |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Alpha-elapitoxin-Dpp2d | 72 | Dendraspis polylepis polylepis | Mutation(s): 0 | 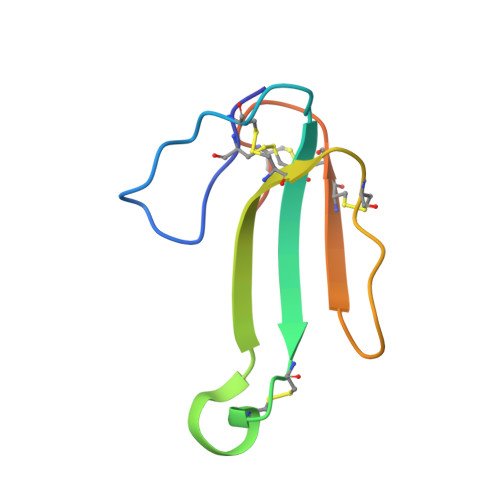 | |
UniProt | |||||
Find proteins for C0HJD7 (Dendroaspis polylepis polylepis) Explore C0HJD7 Go to UniProtKB: C0HJD7 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | C0HJD7 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| GOL Query on GOL | AA [auth C], T [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| ZN Query on ZN | DA [auth D] EA [auth D] FA [auth D] G [auth A] GA [auth D] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| ACT Query on ACT | BA [auth C] CA [auth C] HA [auth D] IA [auth D] N [auth A] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 59.703 | α = 90 |
| b = 145.537 | β = 104.9 |
| c = 64.207 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| SCALEPACK | data scaling |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data reduction |
| BALBES | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Bill & Melinda Gates Foundation | United States | -- |
| National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID) | United States | -- |