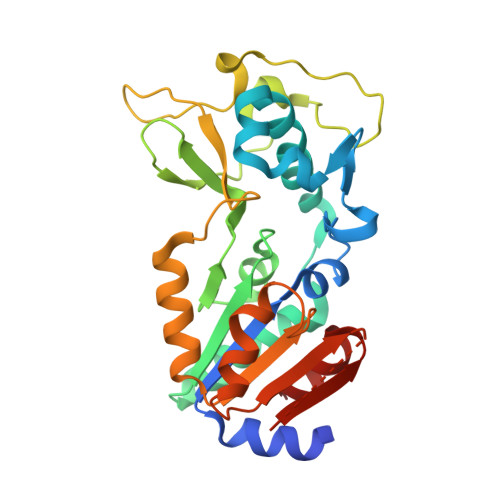
Human SIRT5 variants with reduced stability and activity do not cause neuropathology in mice.
Yuan, T., Kumar, S., Skinner, M.E., Victor-Joseph, R., Abuaita, M., Keijer, J., Zhang, J., Kunkel, T.J., Liu, Y., Petrunak, E.M., Saunders, T.L., Lieberman, A.P., Stuckey, J.A., Neamati, N., Al-Murshedi, F., Alfadhel, M., Spelbrink, J.N., Rodenburg, R., de Boer, V.C.J., Lombard, D.B.(2024) iScience 27: 109991-109991
- PubMed: 38846003
- DOI: https://doi.org/10.1016/j.isci.2024.109991
- Primary Citation of Related Structures:
8GBL, 8GBN - PubMed Abstract:
SIRT5 is a sirtuin deacylase that removes negatively charged lysine modifications, in the mitochondrial matrix and elsewhere in the cell. In benign cells and mouse models, under basal conditions, the phenotypes of SIRT5 deficiency are quite subtle. Here, we identify two homozygous SIRT5 variants in patients suspected to have mitochondrial disease. Both variants, P114T and L128V, are associated with reduced SIRT5 protein stability and impaired biochemical activity, with no evidence of neomorphic or dominant negative properties. The crystal structure of the P114T enzyme was solved and shows only subtle deviations from wild-type. Via CRISPR-Cas9, we generated a mouse model that recapitulates the human P114T mutation; homozygotes show reduced SIRT5 levels and activity, but no obvious metabolic abnormalities, neuropathology, or other gross phenotypes. We conclude that these human SIRT5 variants most likely represent severe hypomorphs, but are likely not by themselves the primary pathogenic cause of the neuropathology observed in the patients.
- Human and Animal Physiology, Wageningen University, De Elst 1, Wageningen, the Netherlands.
Organizational Affiliation: