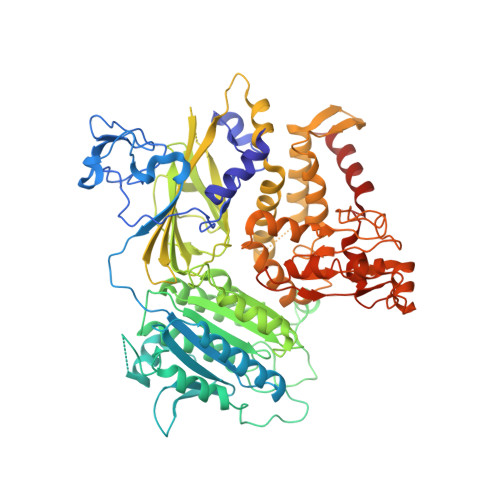
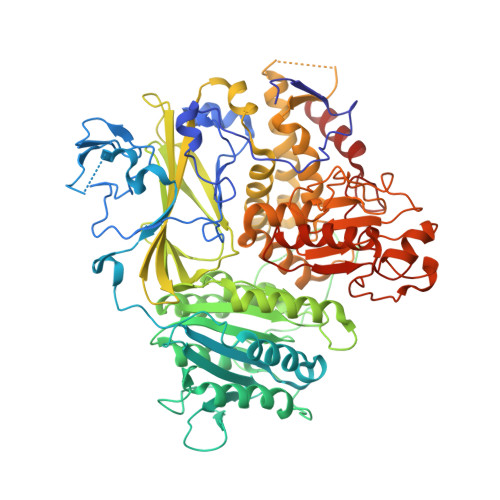
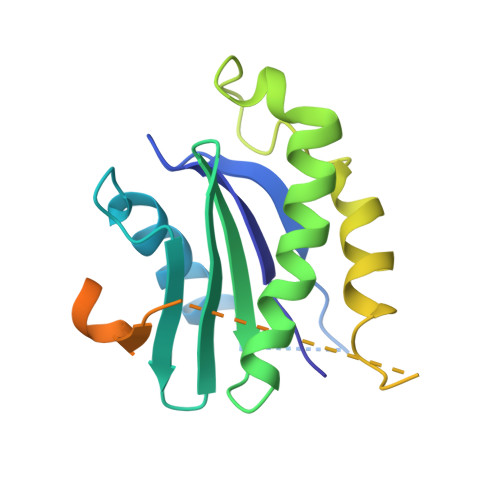
SARS-CoV-2 spike host cell surface exposure promoted by a COPI sorting inhibitor.
Li, Y., Yang, M., Nan, Y., Wang, J., Wang, S., Cui, D., Guo, J., He, P., Dai, W., Zhou, S., Zhang, Y., Ma, W.(2023) Acta Pharm Sin B 13: 3043-3053
- PubMed: 37360012
- DOI: https://doi.org/10.1016/j.apsb.2023.04.007
- Primary Citation of Related Structures:
8HQT, 8HQV, 8HQW, 8HQX, 8HR0 - PubMed Abstract:
Via an insufficient coat protein complex I (COPI) retrieval signal, the majority of SARS-CoV-2 spike (S) is resident in host early secretory organelles and a tiny amount is leaked out in cell surface. Only surface-exposed S can be recognized by B cell receptor (BCR) or anti-S therapeutic monoclonal antibodies (mAbs) that is the trigger step for B cell activation after S mRNA vaccination or infected cell clearance by S mAbs. Now, a drug strategy to promote S host surface exposure is absent. Here, we first combined structural and biochemical analysis to characterize S COPI sorting signals. A potent S COPI sorting inhibitor was then invented, evidently capable of promoting S surface exposure and facilitating infected cell clearance by S antibody-dependent cellular cytotoxicity (ADCC). Importantly, with the inhibitor as a probe, we revealed Omicron BA.1 S is less cell surface exposed than prototypes because of a constellation of S folding mutations, possibly corresponding to its ER chaperone association. Our findings not only suggest COPI is a druggable target against COVID-19, but also highlight SARS-CoV-2 evolution mechanism driven by S folding and trafficking mutations.
- School of Life Science, Beijing University of Chinese Medicine, Beijing 102488, China.
Organizational Affiliation: