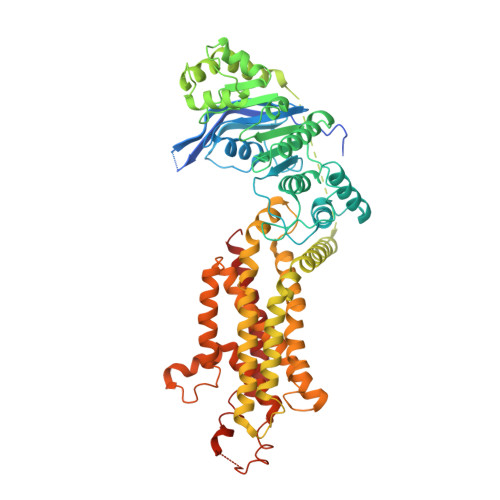
Cryo-EM structure and molecular mechanism of abscisic acid transporter ABCG25.
Huang, X., Zhang, X., An, N., Zhang, M., Ma, M., Yang, Y., Jing, L., Wang, Y., Chen, Z., Zhang, P.(2023) Nat Plants 9: 1709-1719
- PubMed: 37666961
- DOI: https://doi.org/10.1038/s41477-023-01509-7
- Primary Citation of Related Structures:
8I38, 8I39, 8I3A, 8I3B, 8I3C, 8I3D - PubMed Abstract:
Abscisic acid (ABA) is one of the plant hormones that regulate various physiological processes, including stomatal closure, seed germination and development. ABA is synthesized mainly in vascular tissues and transported to distal sites to exert its physiological functions. Many ABA transporters have been identified, however, the molecular mechanism of ABA transport remains elusive. Here we report the cryogenic electron microscopy structure of the Arabidopsis thaliana adenosine triphosphate-binding cassette G subfamily ABA exporter ABCG25 (AtABCG25) in inward-facing apo conformation, ABA-bound pre-translocation conformation and outward-facing occluded conformation. Structural and biochemical analyses reveal that the ABA bound with ABCG25 adopts a similar configuration as that in ABA receptors and that the ABA-specific binding is dictated by residues from transmembrane helices TM1, TM2 and TM5a of each protomer at the transmembrane domain interface. Comparison of different conformational structures reveals conformational changes, especially those of transmembrane helices and residues constituting the substrate translocation pathway during the cross-membrane transport process. Based on the structural data, a 'gate-flipper' translocation model of ABCG25-mediated ABA cross-membrane transport is proposed. Our structural data on AtABCG25 provide new clues to the physiological study of ABA and shed light on its potential applications in plants and agriculture.
- National Key Laboratory of Plant Molecular Genetics, Center for Excellence in Molecular Plant Sciences, Institute of Plant Physiology and Ecology, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: