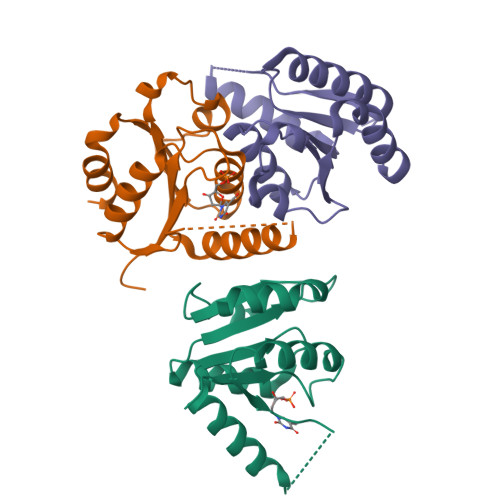
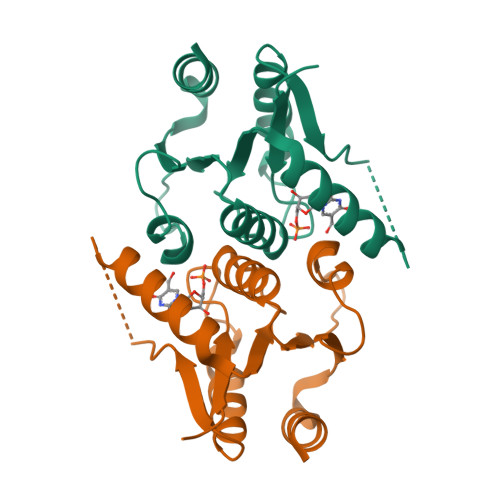
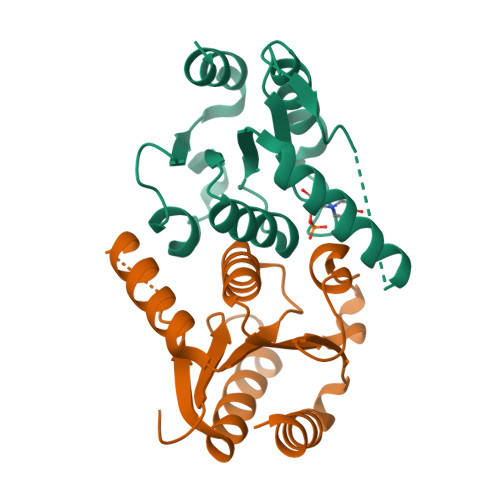
Human 2'-Deoxynucleoside 5'-Phosphate N-Hydrolase 1: The Catalytic Roles of Tyr24 and Asp80.
Carberry, A.E., Devi, S., Harrison, D.J., da Silva, R.G.(2024) Chembiochem 25: e202400047-e202400047
- PubMed: 38350003
- DOI: https://doi.org/10.1002/cbic.202400047
- Primary Citation of Related Structures:
8RPS, 8RPT, 8RQD - PubMed Abstract:
The human enzyme 2'-deoxynucleoside 5'-phosphate N-hydrolase 1 (HsDNPH1) catalyses the hydrolysis of 5-hydroxymethyl-2'-deoxyuridine 5'-phosphate to generate 5-hydroxymethyluracil and 2-deoxyribose-5-phosphate via a covalent 5-phospho-2-deoxyribosylated enzyme intermediate. HsDNPH1 is a promising target for inhibitor development towards anticancer drugs. Here, site-directed mutagenesis of conserved active-site residues, followed by HPLC analysis of the reaction and steady-state kinetics are employed to reveal the importance of each of these residues in catalysis, and the reaction pH-dependence is perturbed by each mutation. Solvent deuterium isotope effects indicate no rate-limiting proton transfers. Crystal structures of D80N-HsDNPH1 in unliganded and substrate-bound states, and of unliganded D80A- and Y24F-HsDNPH1 offer atomic level insights into substrate binding and catalysis. The results reveal a network of hydrogen bonds involving the substrate and the E104-Y24-D80 catalytic triad and are consistent with a proposed mechanism whereby D80 is important for substrate positioning, for helping modulate E104 nucleophilicity, and as the general acid in the first half-reaction. Y24 positions E104 for catalysis and prevents a catalytically disruptive close contact between E104 and D80.
Organizational Affiliation:
School of Biology, Biomedical Sciences Research Complex, University of St Andrews, St Andrews, KY16 9ST, United Kingdom.