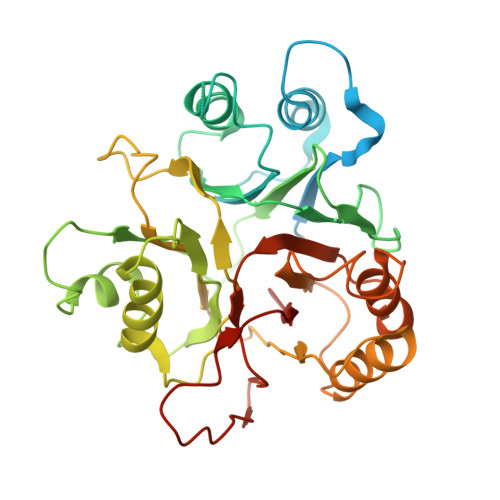
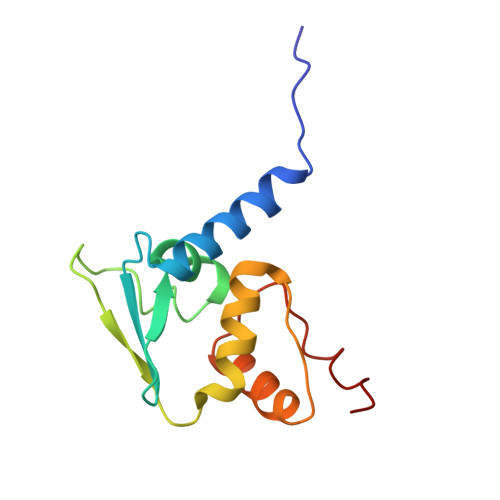
Recognition of BACH1 quaternary structure degrons by two F-box proteins under oxidative stress.
Cao, S., Garcia, S.F., Shi, H., James, E.I., Kito, Y., Shi, H., Mao, H., Kaisari, S., Rona, G., Deng, S., Goldberg, H.V., Ponce, J., Ueberheide, B., Lignitto, L., Guttman, M., Pagano, M., Zheng, N.(2024) Cell 187: 7568-7584.e22
- PubMed: 39504958
- DOI: https://doi.org/10.1016/j.cell.2024.10.012
- Primary Citation of Related Structures:
8UA3, 8UA6, 8UAH, 8UBT, 8UBU, 8UBV - PubMed Abstract:
Ubiquitin-dependent proteolysis regulates diverse cellular functions with high substrate specificity, which hinges on the ability of ubiquitin E3 ligases to decode the targets' degradation signals, i.e., degrons. Here, we show that BACH1, a transcription repressor of antioxidant response genes, features two distinct unconventional degrons encrypted in the quaternary structure of its homodimeric BTB domain. These two degrons are both functionalized by oxidative stress and are deciphered by two complementary E3s. FBXO22 recognizes a degron constructed by the BACH1 BTB domain dimer interface, which is unmasked from transcriptional co-repressors after oxidative stress releases BACH1 from chromatin. When this degron is impaired by oxidation, a second BACH1 degron manifested by its destabilized BTB dimer is probed by a pair of FBXL17 proteins that remodels the substrate into E3-bound monomers for ubiquitination. Our findings highlight the multidimensionality of protein degradation signals and the functional complementarity of different ubiquitin ligases targeting the same substrate.
- Department of Pharmacology, University of Washington, Box 357280, Seattle, WA 98195, USA; Howard Hughes Medical Institute, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: