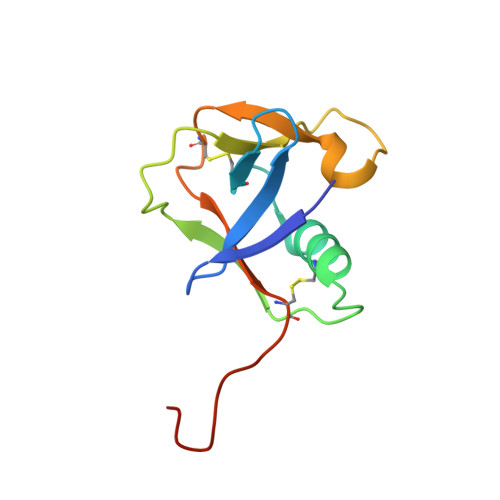
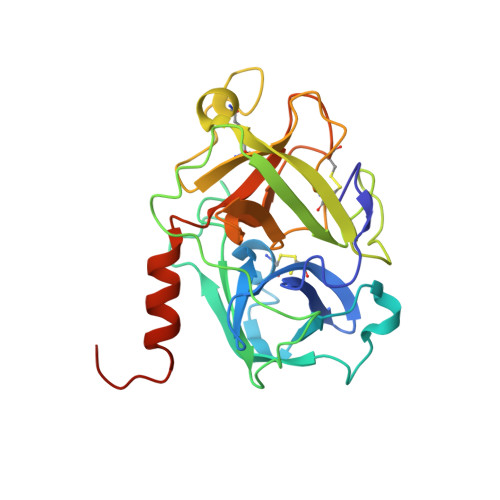
TMPRSS2 complexed with the noncovalent inhibitor 6-amidino-2-napthol
Fraser, B.J., Dong, A., Kutera, M., Seitova, A., Li, Y., Hutchinson, A., Edwards, A., Benard, F., Halabelian, L., Arrowsmith, C., Structural Genomics Consortium (SGC)To be published.