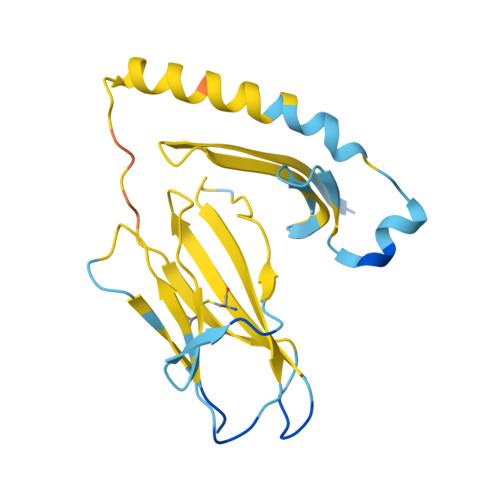
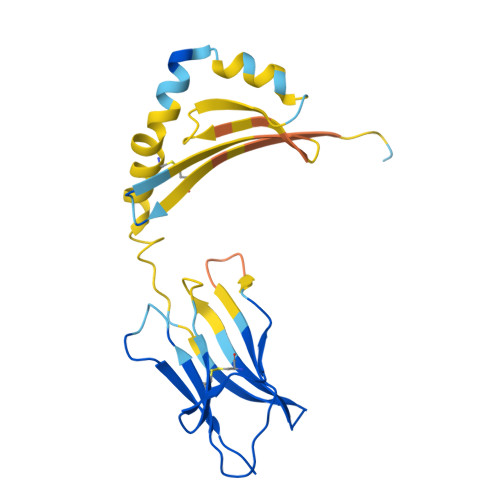
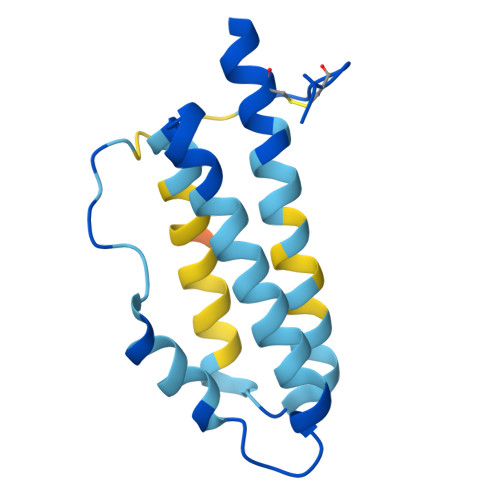
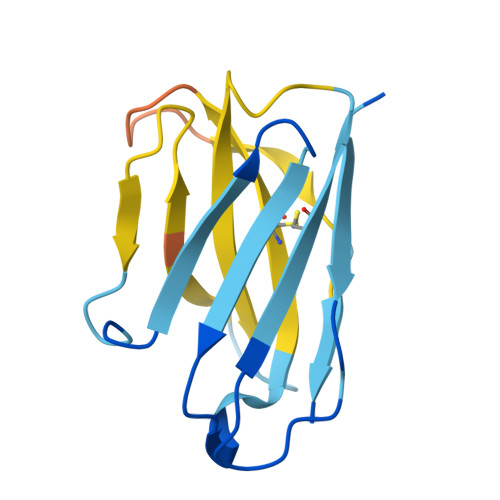
A general system for targeting MHC class II-antigen complex via a single adaptable loop.
Du, H., Liu, J., Jude, K.M., Yang, X., Li, Y., Bell, B., Yang, H., Kassardjian, A., Blackson, W., Mobedi, A., Parekh, U., Parra Sperberg, R.A., Julien, J.P., Mellins, E.D., Garcia, K.C., Huang, P.S.(2024) Nat Biotechnol
- PubMed: 39672953
- DOI: https://doi.org/10.1038/s41587-024-02466-y
- Primary Citation of Related Structures:
8VSJ - PubMed Abstract:
Major histocompatibility complex class II (MHCII) bound to a peptide antigen mediates interactions between CD4 + T cells and antigen-presenting cells. Targeting peptide-MHCII with T cell antigen receptors (TCRs) and TCR-like antibodies has shown promise for autoimmune diseases and microbiome tolerance. To develop a general targeting approach, we introduce targeted recognition of antigen-MHC complex reporter for MHCII (TRACeR-II) for the rapid development of peptide-specific MHCII binders. TRACeR-II binders have a small helical bundle scaffold and use a single loop to recognize peptide-MHCII, which offers versatility and enables structural modeling of the interactions to target MHCII antigens. We demonstrate rapid generation of TRACeR-II binders to multiple molecules with affinities in the low-nanomolar to low-micromolar range, comparable to best-in-class TCRs and antibodies. Through computational protein design, we created specific binding sequences in silico from only the sequence of a severe acute respiratory syndrome coronavirus 2 peptide. TRACeR-II provides a straightforward approach to target antigen-MHCII without relying on combinatorial selection on complementarity-determining region loops.
Organizational Affiliation:
Department of Chemistry, Stanford University, Stanford, CA, USA.