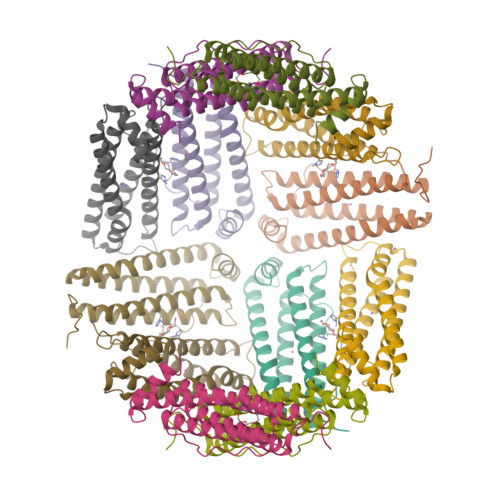
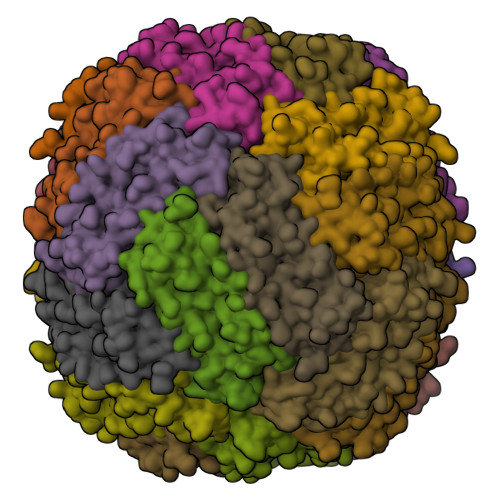
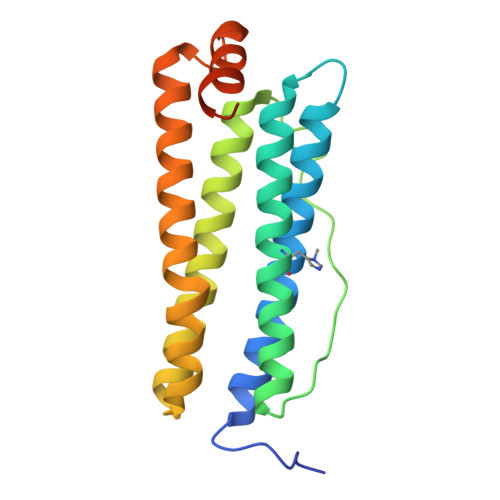
Site-Specific Histidine Aza-Michael Addition in Proteins Enabled by a Ferritin-Based Metalloenzyme.
Tsou, J.C., Tsou, C.J., Wang, C.H., Ko, A.A., Wang, Y.H., Liang, H.H., Sun, J.C., Huang, K.F., Ko, T.P., Lin, S.Y., Wang, Y.S.(2024) J Am Chem Soc 146: 33309-33315
- PubMed: 39499210
- DOI: https://doi.org/10.1021/jacs.4c14446
- Primary Citation of Related Structures:
9JIU, 9JQB, 9JQC, 9JQD, 9JQE - PubMed Abstract:
Histidine modifications of proteins are broadly based on chemical methods triggering N-substitution reactions such as aza-Michael addition at histidine's moderately nucleophilic imidazole side chain. While recent studies have demonstrated chemoselective, histidine-specific modifications by further exploiting imidazole's electrophilic reactivity to overcome interference from the more nucleophilic lysine and cysteine, achieving site-specific histidine modifications remains a major challenge due to the absence of spatial control over chemical processes. Herein, through X-ray crystallography and cryo-electron microscopy structural studies, we describe the rational design of a nature-inspired, noncanonical amino-acid-incorporated, human ferritin-based metalloenzyme that is capable of introducing site-specific post-translational modifications (PTMs) to histidine in peptides and proteins. Specifically, chemoenzymatic aza-Michael additions on single histidine residues were carried out on eight protein substrates ranging from 10 to 607 amino acids including the insulin peptide hormone. By introducing an insulin-targeting peptide into our metalloenzyme, we further directed modifications to be carried out site-specifically on insulin's B-chain histidine 5. The success of this biocatalysis platform outlines a novel approach in introducing residue- and, moreover, site-specific post-translational modifications to peptides and proteins, which may further enable reactions to be carried out in vivo .
Organizational Affiliation:
Institute of Biological Chemistry, Academia Sinica, Taipei 11529, Taiwan.