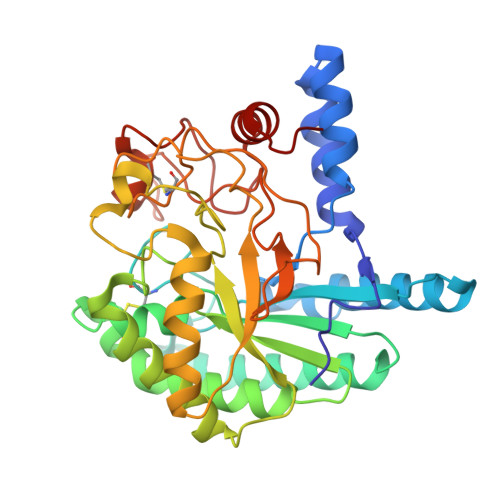
Structural Basis for Ligand Binding and Processivity in Cellobiohydrolase Cel6A from Humicola Insolens
Varrot, A., Frandsen, T.P., Von Ossowski, I., Boyer, V., Driguez, H., Schulein, M., Davies, G.J.(2003) Structure 11: 855
- PubMed: 12842048
- DOI: https://doi.org/10.1016/s0969-2126(03)00124-2
- Primary Citation of Related Structures:
1OC5, 1OC6, 1OC7, 1OCB, 1OCJ - PubMed Abstract:
The enzymatic digestion of cellulose entails intimate involvement of cellobiohydrolases, whose characteristic active-center tunnel contributes to a processive degradation of the polysaccharide. The cellobiohydrolase Cel6A displays an active site within a tunnel formed by two extended loops, which are known to open and close in response to ligand binding. Here we present five structures of wild-type and mutant forms of Cel6A from Humicola insolens in complex with nonhydrolyzable thio-oligosaccharides, at resolutions from 1.7-1.1 A, dissecting the structural accommodation of a processing substrate chain through the active center during hydrolysis. Movement of ligand is facilitated by extensive solvent-mediated interactions and through flexibility in the hydrophobic surfaces provided by a sheath of tryptophan residues.
- Department of Chemistry, University of York, Heslington, Y010 5YW, York, United Kingdom.
Organizational Affiliation: