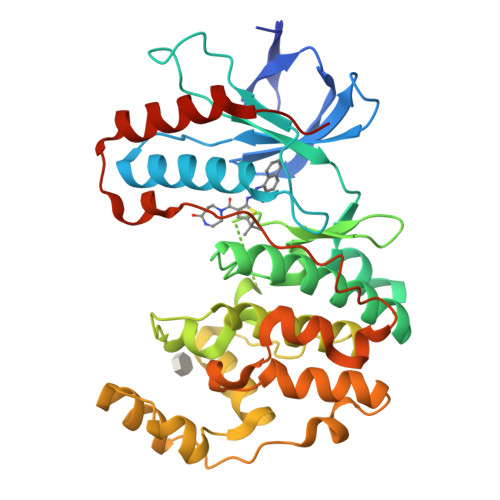
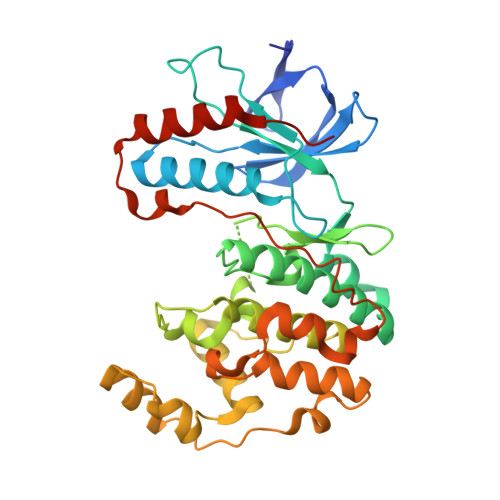
Discovery of a novel class of non-ATP site DFG-out state p38 inhibitors utilizing computationally assisted virtual fragment-based drug design (vFBDD).
Moffett, K., Konteatis, Z., Nguyen, D., Shetty, R., Ludington, J., Fujimoto, T., Lee, K.J., Chai, X., Namboodiri, H., Karpusas, M., Dorsey, B., Guarnieri, F., Bukhtiyarova, M., Springman, E., Michelotti, E.(2011) Bioorg Med Chem Lett 21: 7155-7165
- PubMed: 22014550
- DOI: https://doi.org/10.1016/j.bmcl.2011.09.078
- Primary Citation of Related Structures:
3P5K, 3P78, 3P79, 3P7A, 3P7B, 3P7C - PubMed Abstract:
Discovery of a new class of DFG-out p38α kinase inhibitors with no hinge interaction is described. A computationally assisted, virtual fragment-based drug design (vFBDD) platform was utilized to identify novel non-aromatic fragments which make productive hydrogen bond interactions with Arg 70 on the αC-helix. Molecules incorporating these fragments were found to be potent inhibitors of p38 kinase. X-ray co-crystal structures confirmed the predicted binding modes. A lead compound was identified as a potent (p38α IC(50)=22 nM) and highly selective (≥ 150-fold against 150 kinase panel) DFG-out p38 kinase inhibitor.
Organizational Affiliation:
Ansaris, Four Valley Square, 512 East Township Line Road, Blue Bell, PA 19422, USA. kmistry@earthlink.net