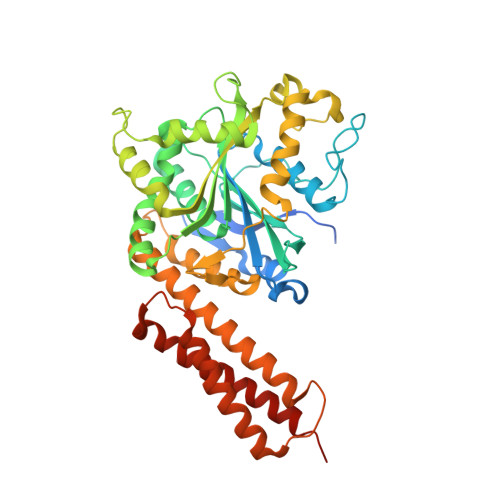
Structural basis for the nucleotide-dependent dimerization of the large G protein atlastin-1/SPG3A.
Byrnes, L.J., Sondermann, H.(2011) Proc Natl Acad Sci U S A 108: 2216-2221
- PubMed: 21220294
- DOI: https://doi.org/10.1073/pnas.1012792108
- Primary Citation of Related Structures:
3Q5D, 3Q5E - PubMed Abstract:
The large GTPase atlastin belongs to the dynamin superfamily that has been widely implicated in facilitating membrane tubulation, fission, and in select cases, fusion. Mutations spread across atlastin isoform 1 (atlastin-1) have been identified in patients suffering from hereditary spastic paraplegia (HSP), a neurodegenerative disorder affecting motor neuron function in the lower extremities. On a molecular level, atlastin-1 associates with high membrane curvature and fusion events at the endoplasmic reticulum and cis-Golgi. Here we report crystal structures of atlastin-1 comprising the G and middle domains in two different conformations. Although the orientation of the middle domain relative to the G domain is different in the two structures, both reveal dimeric assemblies with a common, GDP-bound G domain dimer. In contrast, dimer formation in solution is observed only in the presence of GTP and transition state analogs, similar to other G proteins that are activated by nucleotide-dependent dimerization. Analyses of solution scattering data suggest that upon nucleotide binding, the protein adopts a somewhat extended, dimeric conformation that is reminiscent of one of the two crystal structures. These structural studies suggest a model for nucleotide-dependent regulation of atlastin with implications for membrane fusion. This mechanism is affected in several mutants associated with HSP, providing insights into disease pathogenesis.
- Department of Molecular Medicine, College of Veterinary Medicine, Cornell University, Ithaca, NY 14853, USA.
Organizational Affiliation: