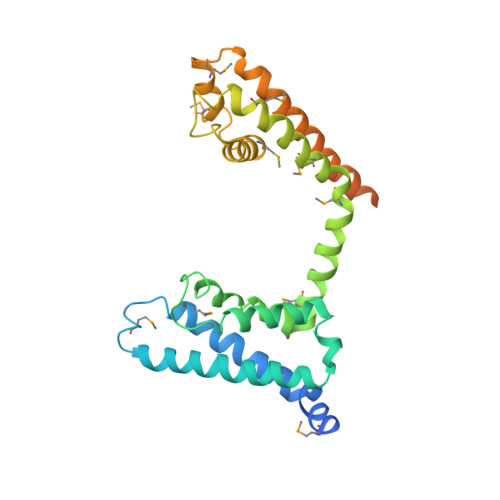
Crystal structure of a voltage-gated sodium channel in two potentially inactivated states.
Payandeh, J., Gamal El-Din, T.M., Scheuer, T., Zheng, N., Catterall, W.A.(2012) Nature 486: 135-139
- PubMed: 22678296
- DOI: https://doi.org/10.1038/nature11077
- Primary Citation of Related Structures:
4EKW - PubMed Abstract:
In excitable cells, voltage-gated sodium (Na(V)) channels activate to initiate action potentials and then undergo fast and slow inactivation processes that terminate their ionic conductance. Inactivation is a hallmark of Na(V) channel function and is critical for control of membrane excitability, but the structural basis for this process has remained elusive. Here we report crystallographic snapshots of the wild-type Na(V)Ab channel from Arcobacter butzleri captured in two potentially inactivated states at 3.2 Å resolution. Compared to previous structures of Na(V)Ab channels with cysteine mutations in the pore-lining S6 helices (ref. 4), the S6 helices and the intracellular activation gate have undergone significant rearrangements: one pair of S6 helices has collapsed towards the central pore axis and the other S6 pair has moved outward to produce a striking dimer-of-dimers configuration. An increase in global structural asymmetry is observed throughout our wild-type Na(V)Ab models, reshaping the ion selectivity filter at the extracellular end of the pore, the central cavity and its residues that are analogous to the mammalian drug receptor site, and the lateral pore fenestrations. The voltage-sensing domains have also shifted around the perimeter of the pore module in wild-type Na(V)Ab, compared to the mutant channel, and local structural changes identify a conserved interaction network that connects distant molecular determinants involved in Na(V) channel gating and inactivation. These potential inactivated-state structures provide new insights into Na(V) channel gating and novel avenues to drug development and therapy for a range of debilitating Na(V) channelopathies.
Organizational Affiliation:
Department of Pharmacology, University of Washington, Seattle, Washington 98195, USA.