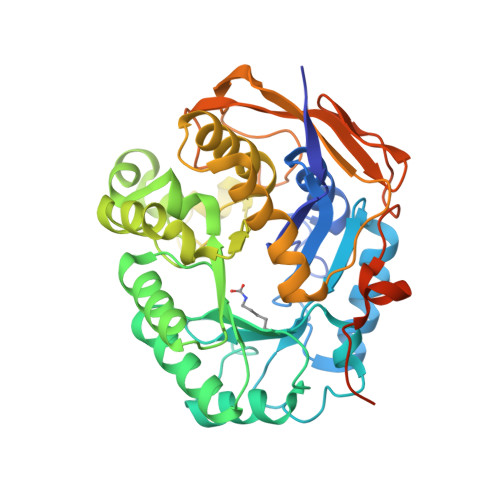
Characterization of the catalytic flexible loop in the dihydroorotase domain of the human multi-enzymatic protein CAD.
Del Cano-Ochoa, F., Grande-Garcia, A., Reverte-Lopez, M., D'Abramo, M., Ramon-Maiques, S.(2018) J Biological Chem 293: 18903-18913
- PubMed: 30315107
- DOI: https://doi.org/10.1074/jbc.RA118.005494
- Primary Citation of Related Structures:
6HFD, 6HFE, 6HFF, 6HFH, 6HFI, 6HFJ, 6HFK, 6HFL, 6HFN, 6HFP, 6HFQ, 6HFR, 6HFS, 6HFU, 6HG1, 6HG2, 6HG3 - PubMed Abstract:
The dihydroorotase (DHOase) domain of the multifunctional protein carbamoyl-phosphate synthetase 2, aspartate transcarbamoylase, and dihydroorotase (CAD) catalyzes the third step in the de novo biosynthesis of pyrimidine nucleotides in animals. The crystal structure of the DHOase domain of human CAD (huDHOase) revealed that, despite evolutionary divergence, its active site components are highly conserved with those in bacterial DHOases, encoded as monofunctional enzymes. An important element for catalysis, conserved from Escherichia coli to humans, is a flexible loop that closes as a lid over the active site. Here, we combined mutagenic, structural, biochemical, and molecular dynamics analyses to characterize the function of the flexible loop in the activity of CAD's DHOase domain. A huDHOase chimera bearing the E. coli DHOase flexible loop was inactive, suggesting the presence of distinctive elements in the flexible loop of huDHOase that cannot be replaced by the bacterial sequence. We pinpointed Phe-1563, a residue absolutely conserved at the tip of the flexible loop in CAD's DHOase domain, as a critical element for the conformational equilibrium between the two catalytic states of the protein. Substitutions of Phe-1563 with Ala, Leu, or Thr prevented the closure of the flexible loop and inactivated the protein, whereas substitution with Tyr enhanced the interactions of the loop in the closed position and reduced fluctuations and the reaction rate. Our results confirm the importance of the flexible loop in CAD's DHOase domain and explain the key role of Phe-1563 in configuring the active site and in promoting substrate strain and catalysis.
- From the Department of Genome Dynamics and Function, Centro de Biología Molecular Severo Ochoa (CSIC-UAM), Madrid 28049, Spain.
Organizational Affiliation: