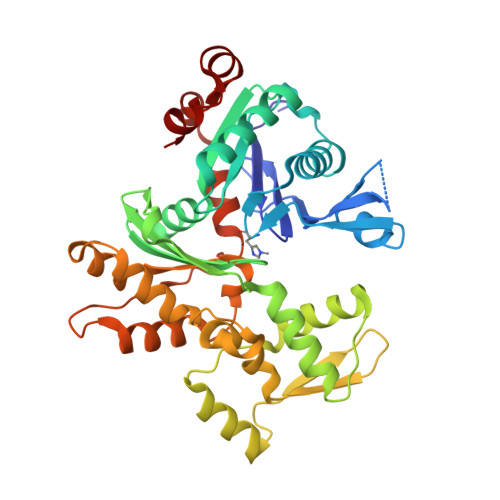
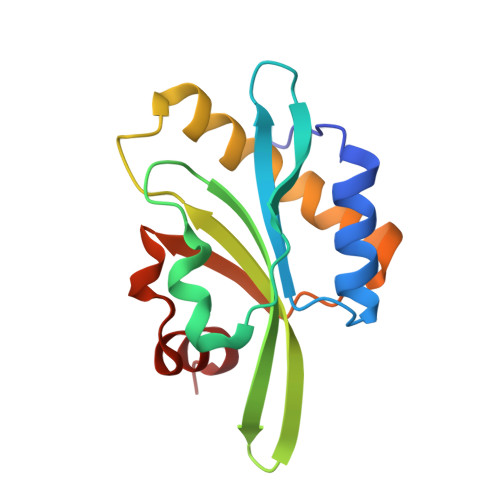
A functional family of fluorescent nucleotide analogues to investigate actin dynamics and energetics.
Colombo, J., Antkowiak, A., Kogan, K., Kotila, T., Elliott, J., Guillotin, A., Lappalainen, P., Michelot, A.(2021) Nat Commun 12: 548-548
- PubMed: 33483497
- DOI: https://doi.org/10.1038/s41467-020-20827-4
- Primary Citation of Related Structures:
6YP9 - PubMed Abstract:
Actin polymerization provides force for vital processes of the eukaryotic cell, but our understanding of actin dynamics and energetics remains limited due to the lack of high-quality probes. Most current probes affect dynamics of actin or its interactions with actin-binding proteins (ABPs), and cannot track the bound nucleotide. Here, we identify a family of highly sensitive fluorescent nucleotide analogues structurally compatible with actin. We demonstrate that these fluorescent nucleotides bind to actin, maintain functional interactions with a number of essential ABPs, are hydrolyzed within actin filaments, and provide energy to power actin-based processes. These probes also enable monitoring actin assembly and nucleotide exchange with single-molecule microscopy and fluorescence anisotropy kinetics, therefore providing robust and highly versatile tools to study actin dynamics and functions of ABPs.
- Aix Marseille Univ, CNRS, IBDM, Turing Centre for Living Systems, 13288, Marseille, France.
Organizational Affiliation: