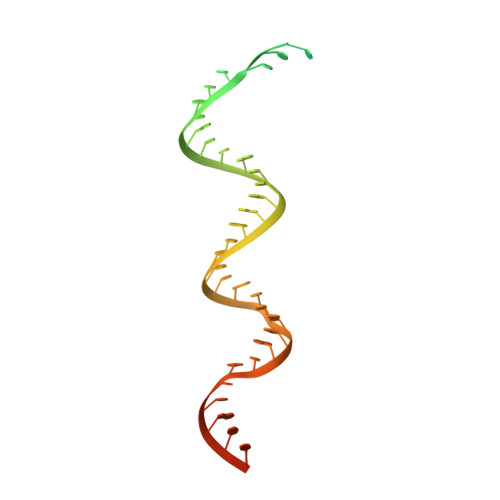
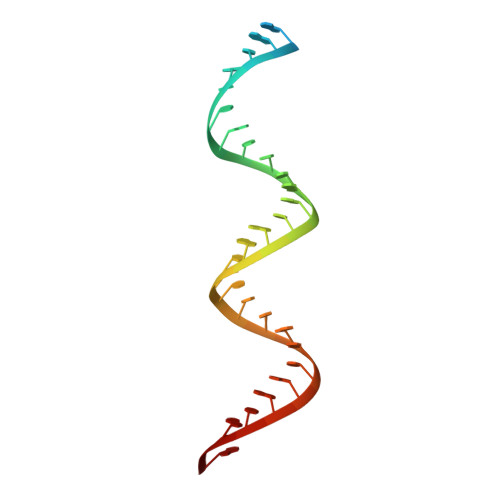
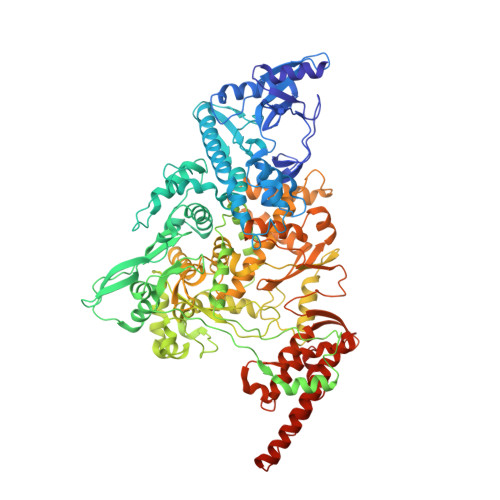
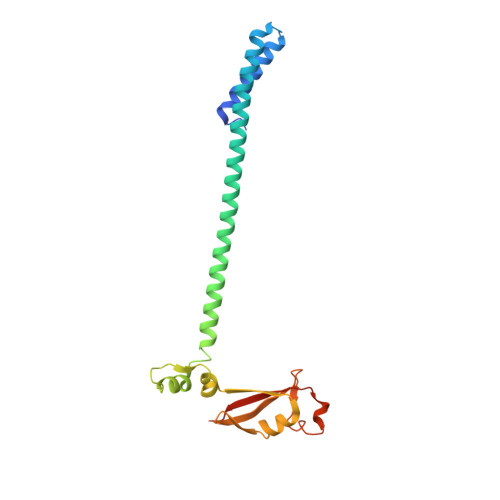
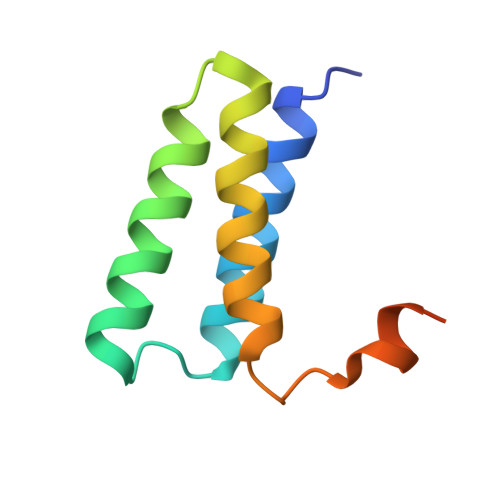
Remdesivir overcomes the S861 roadblock in SARS-CoV-2 polymerase elongation complex.
Wu, J., Wang, H., Liu, Q., Li, R., Gao, Y., Fang, X., Zhong, Y., Wang, M., Wang, Q., Rao, Z., Gong, P.(2021) Cell Rep 37: 109882-109882
- PubMed: 34653416
- DOI: https://doi.org/10.1016/j.celrep.2021.109882
- Primary Citation of Related Structures:
7DTE - PubMed Abstract:
Remdesivir (RDV), a nucleotide analog with broad-spectrum features, has exhibited effectiveness in COVID-19 treatment. However, the precise working mechanism of RDV when targeting the viral RNA-dependent RNA polymerase (RdRP) has not been fully elucidated. Here, we solve a 3.0-Å structure of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) RdRP elongation complex (EC) and assess RDV intervention in polymerase elongation phase. Although RDV could induce an "i+3" delayed termination in meta-stable complexes, only pausing and subsequent elongation are observed in the EC. A comparative investigation using an enterovirus RdRP further confirms similar delayed intervention and demonstrates that steric hindrance of the RDV-characteristic 1'-cyano at the -4 position is responsible for the "i+3" intervention, although two representative Flaviviridae RdRPs do not exhibit similar behavior. A comparison of representative viral RdRP catalytic complex structures indicates that the product RNA backbone encounters highly conserved structural elements, highlighting the broad-spectrum intervention potential of 1'-modified nucleotide analogs in anti-RNA virus drug development.
Organizational Affiliation:
Key Laboratory of Special Pathogens and Biosafety, Wuhan Institute of Virology, Center for Biosafety Mega-Science, Chinese Academy of Sciences, No. 44 Xiao Hong Shan, Wuhan, Hubei 430071, China.