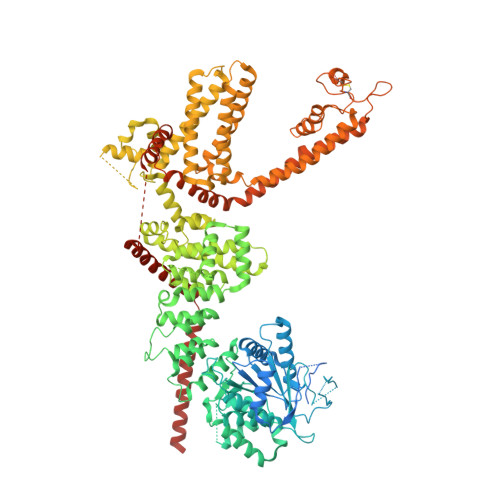
Structures of a mammalian TRPM8 in closed state.
Zhao, C., Xie, Y., Xu, L., Ye, F., Xu, X., Yang, W., Yang, F., Guo, J.(2022) Nat Commun 13: 3113-3113
- PubMed: 35662242
- DOI: https://doi.org/10.1038/s41467-022-30919-y
- Primary Citation of Related Structures:
7WRA, 7WRB, 7WRC, 7WRD, 7WRE, 7WRF - PubMed Abstract:
Transient receptor potential melastatin 8 (TRPM8) channel is a Ca 2+ -permeable non-selective cation channel that acts as the primary cold sensor in humans. TRPM8 is also activated by ligands such as menthol, icilin, and phosphatidylinositol 4,5-bisphosphate (PIP 2 ), and desensitized by Ca 2+ . Here we have determined electron cryo-microscopy structures of mouse TRPM8 in the absence of ligand, and in the presence of Ca 2+ and icilin at 2.5-3.2 Å resolution. The ligand-free state TRPM8 structure represents the full-length structure of mammalian TRPM8 channels with a canonical S4-S5 linker and the clearly resolved selectivity filter and outer pore loop. TRPM8 has a short but wide selectivity filter which may account for its permeability to hydrated Ca 2+ . Ca 2+ and icilin bind in the cytosolic-facing cavity of the voltage-sensing-like domain of TRPM8 but induce little conformational change. All the ligand-bound TRPM8 structures adopt the same closed conformation as the ligand-free structure. This study reveals the overall architecture of mouse TRPM8 and the structural basis for its ligand recognition.
Organizational Affiliation:
Department of Biophysics, and Department of Neurology of the Fourth Affiliated Hospital, Zhejiang University School of Medicine, Hangzhou, Zhejiang, 310058, China.