Structure of human MAIT A-F7 TCR in complex with human MR1-3,4-dihydroxybenzaldehyde
Awad, W., Rossjohn, J.(2024) J Exp Med
Experimental Data Snapshot
Starting Model: experimental
View more details
(2024) J Exp Med
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Major histocompatibility complex class I-related gene protein | 271 | Homo sapiens | Mutation(s): 0 Gene Names: MR1 | 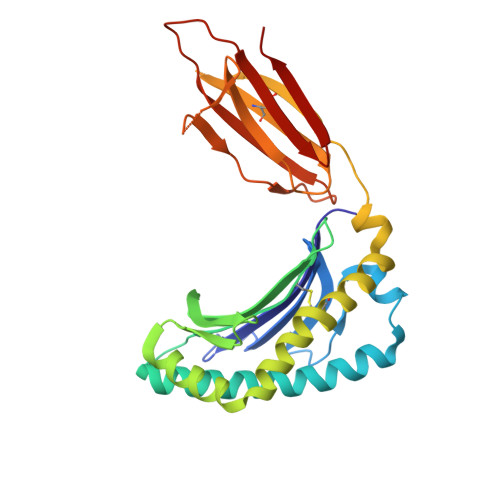 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q95460 (Homo sapiens) Explore Q95460 Go to UniProtKB: Q95460 | |||||
PHAROS: Q95460 GTEx: ENSG00000153029 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q95460 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Beta-2-microglobulin | 100 | Homo sapiens | Mutation(s): 0 Gene Names: B2M, CDABP0092, HDCMA22P | 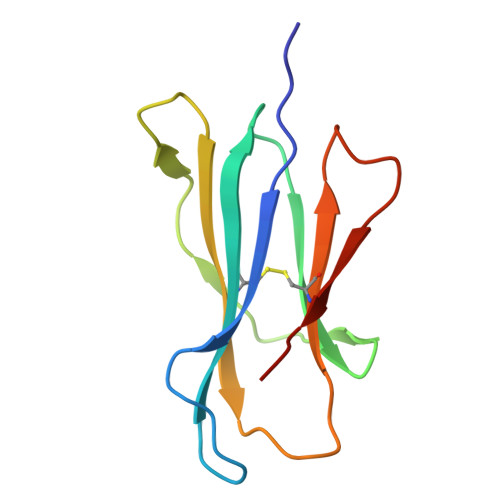 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P61769 (Homo sapiens) Explore P61769 Go to UniProtKB: P61769 | |||||
PHAROS: P61769 GTEx: ENSG00000166710 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P61769 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Human TCR TRAV1-2_ALPHA | 204 | Homo sapiens | Mutation(s): 0 | 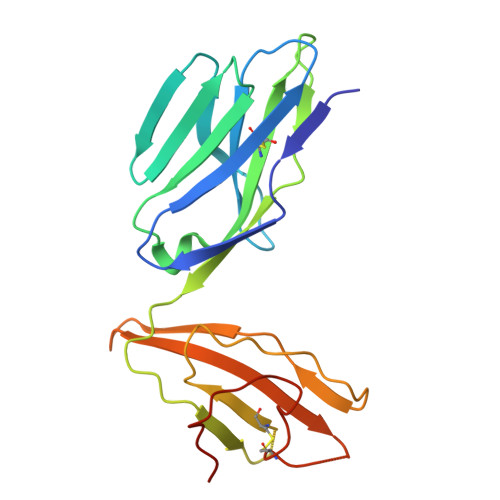 | |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Human TCR TRBV6-1_BETA | 246 | Homo sapiens | Mutation(s): 0 | 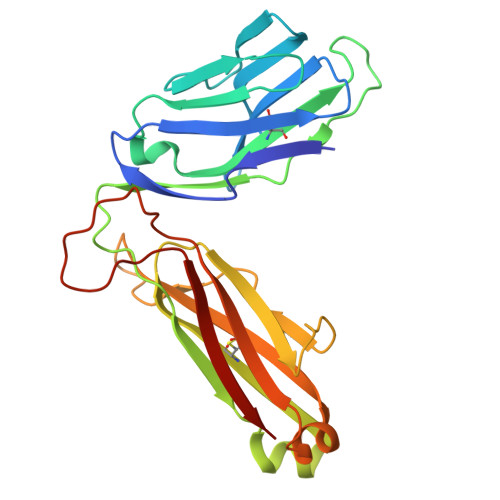 | |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| H6N (Subject of Investigation/LOI) Query on H6N | K [auth A], R [auth C] | Protocatechuic aldehyde C7 H6 O3 IBGBGRVKPALMCQ-UHFFFAOYSA-N |  | ||
| GOL Query on GOL | I [auth A] J [auth A] P [auth A] Q [auth C] V [auth F] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| ACT Query on ACT | L [auth A] M [auth A] N [auth A] O [auth A] S [auth C] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| NA Query on NA | W [auth F], Y [auth H] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 217.129 | α = 90 |
| b = 70.405 | β = 104.61 |
| c = 143.573 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| XDS | data scaling |
| PHENIX | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Australian Research Council (ARC) | Australia | DE220101491 |
| National Health and Medical Research Council (NHMRC, Australia) | Australia | -- |